-
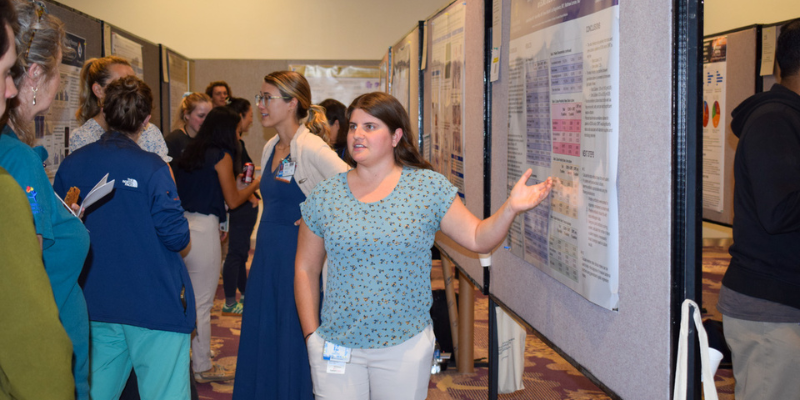

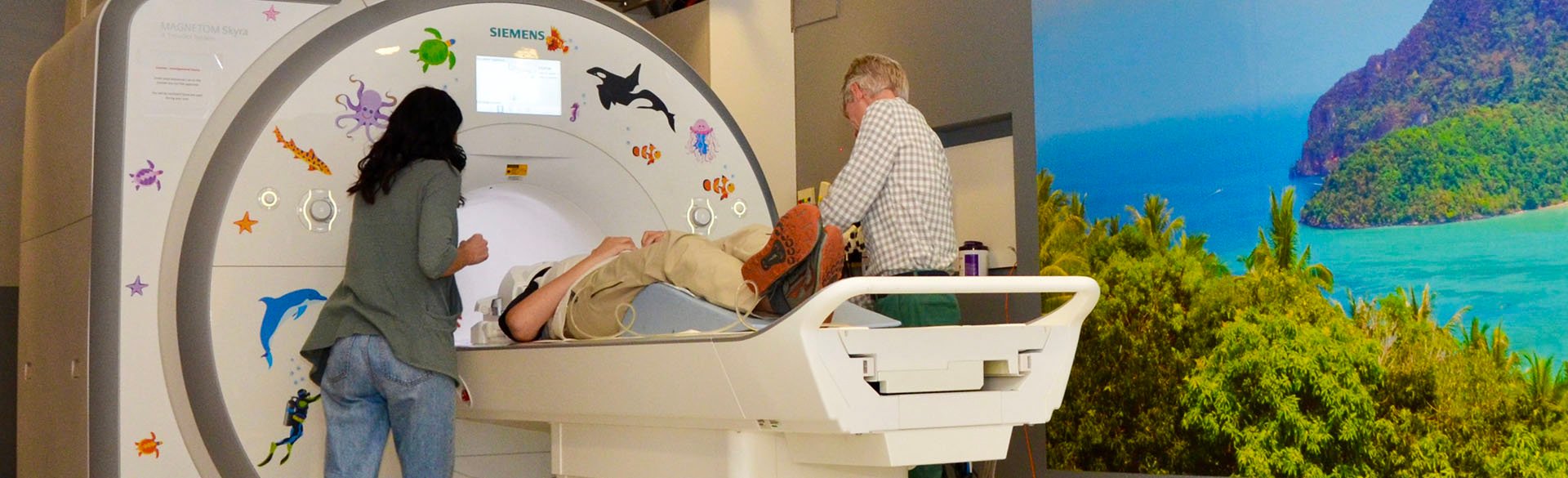
Poster Sessions: a View into the Future of Pediatric Medicine
7/7/2025Discover how the Colorado Clinical and Translational Sciences Institute's Pediatric Research Poster Session fosters innovation and advances child health research and patient care....Read More
-
-1.png)
Director's Corner
6/23/2025Read the latest news from the Colorado Clinical and Translational Sciences Institute (CCTSI)....Read More
-
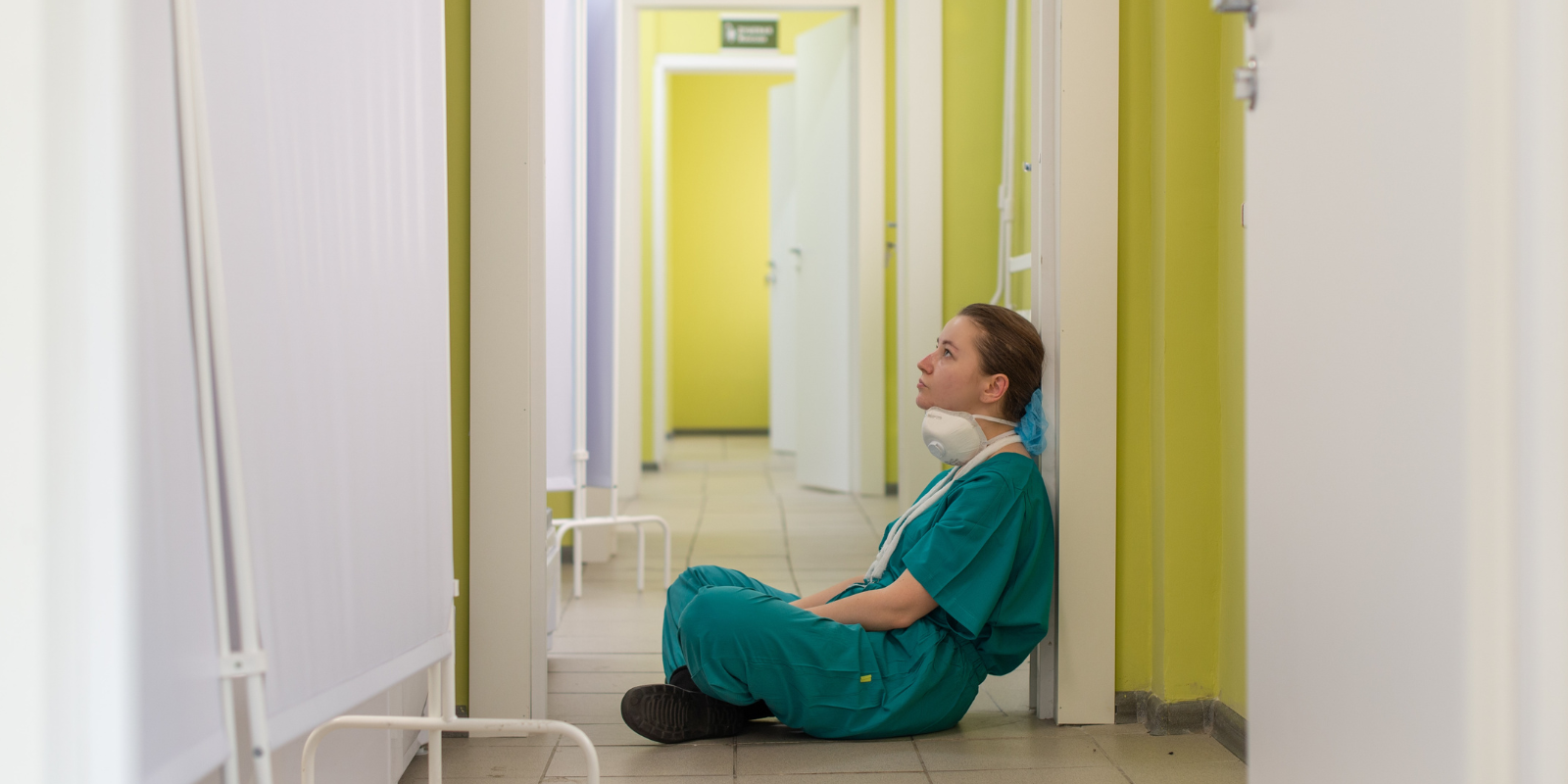
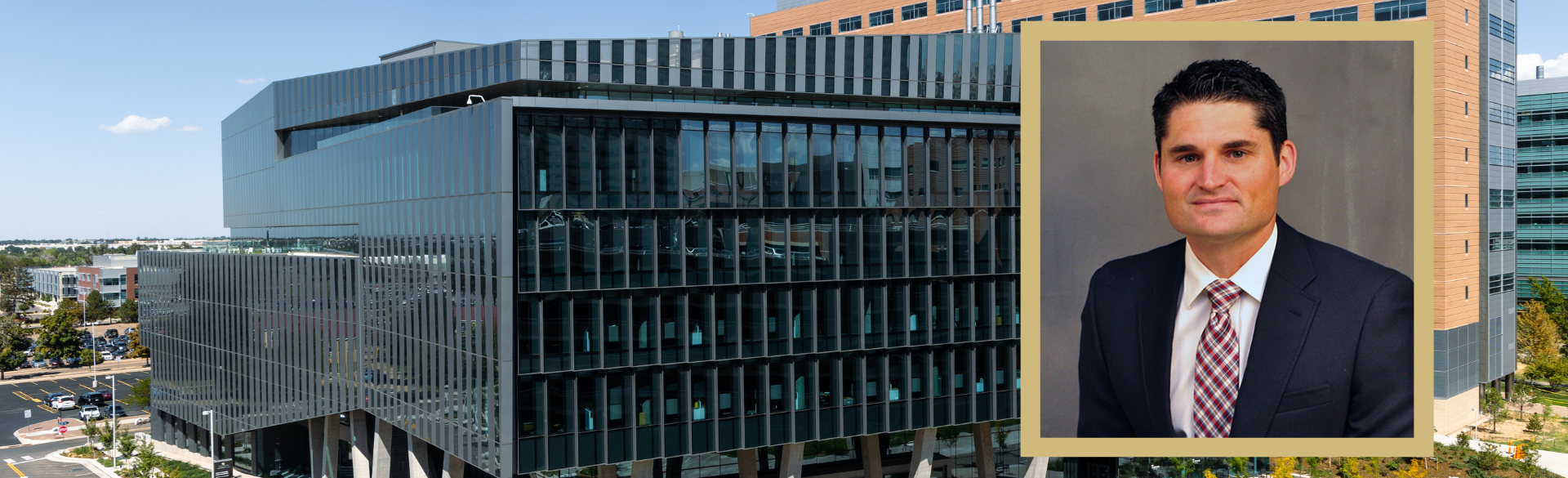
OCCULT-O2: Seeking Ways to Improve Care for Critically Ill Patients on Mechanical Ventilation
6/16/2025Neil Aggarwal, MD, of the CU Department of Medicine is leading a "pragmatic clinical trial" to determine how much oxygen to give patients in the ICU....Read More
-
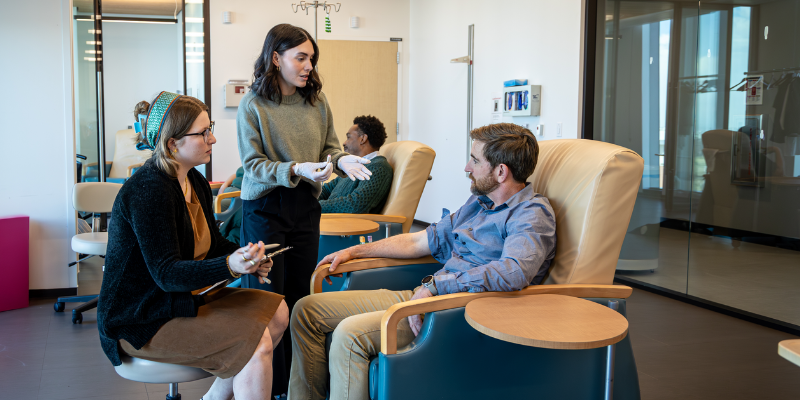

Science, Discovery, Health and Healing
5/5/2025The CCTSI published an annual report highlighting their efforts in 2024....Read More
-
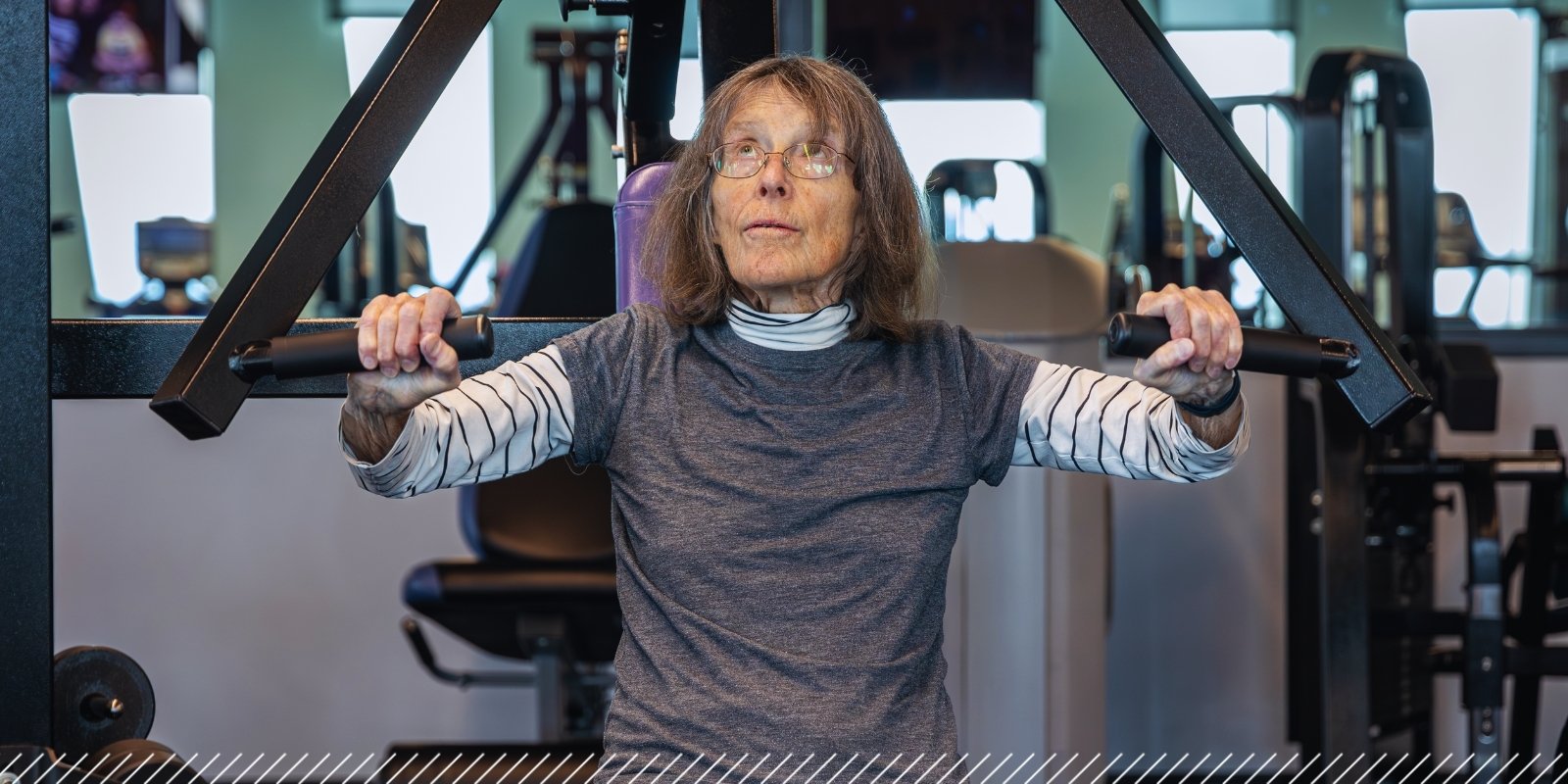

Science Supports Exercise as Medicine; Patients With Parkinson’s See Benefits
5/2/2025Research at the CU Anschutz campus continues to build on the evidence that exercise is the best medicine patients with Parkinson's disease....Read More
-
.png)
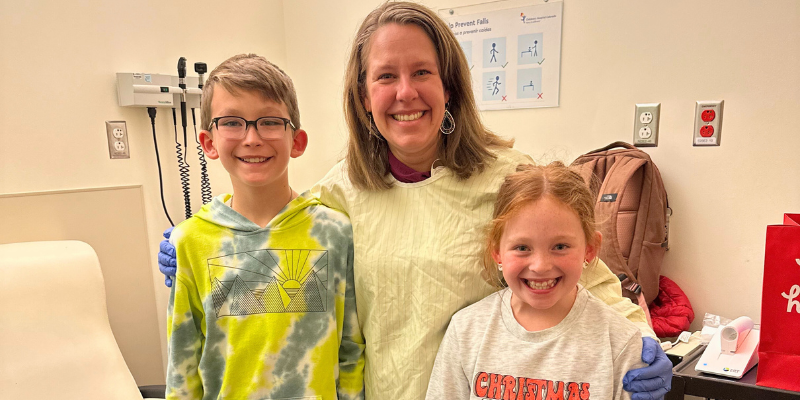
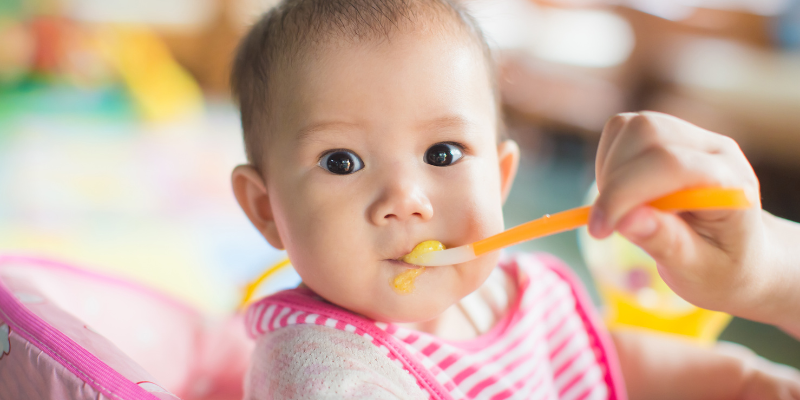
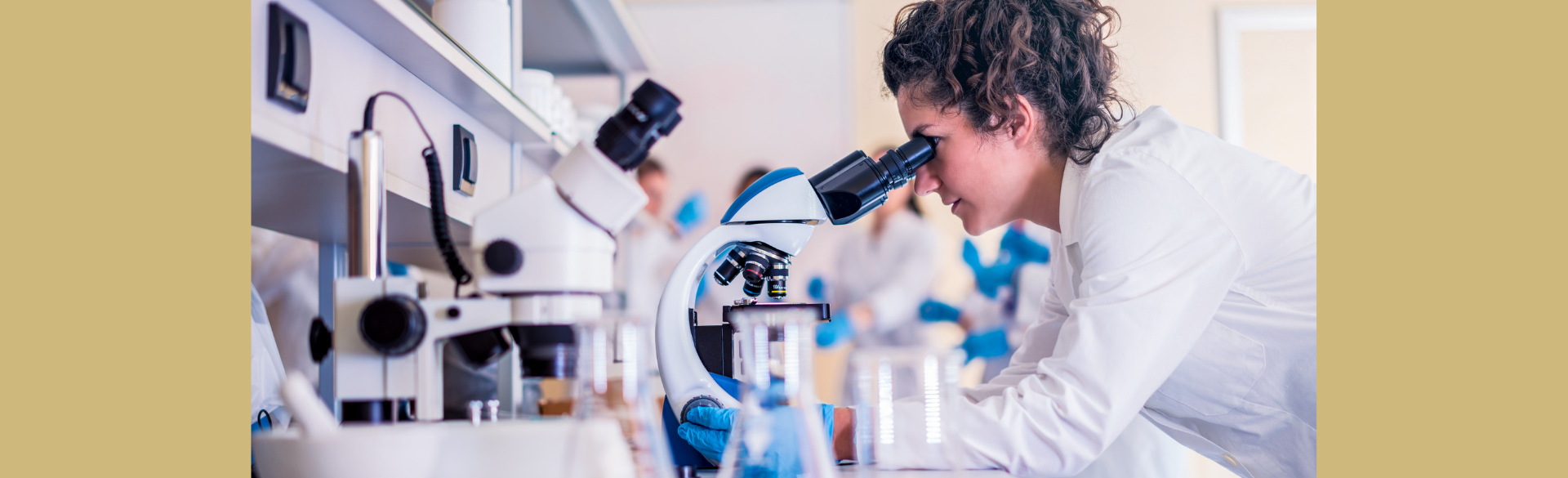
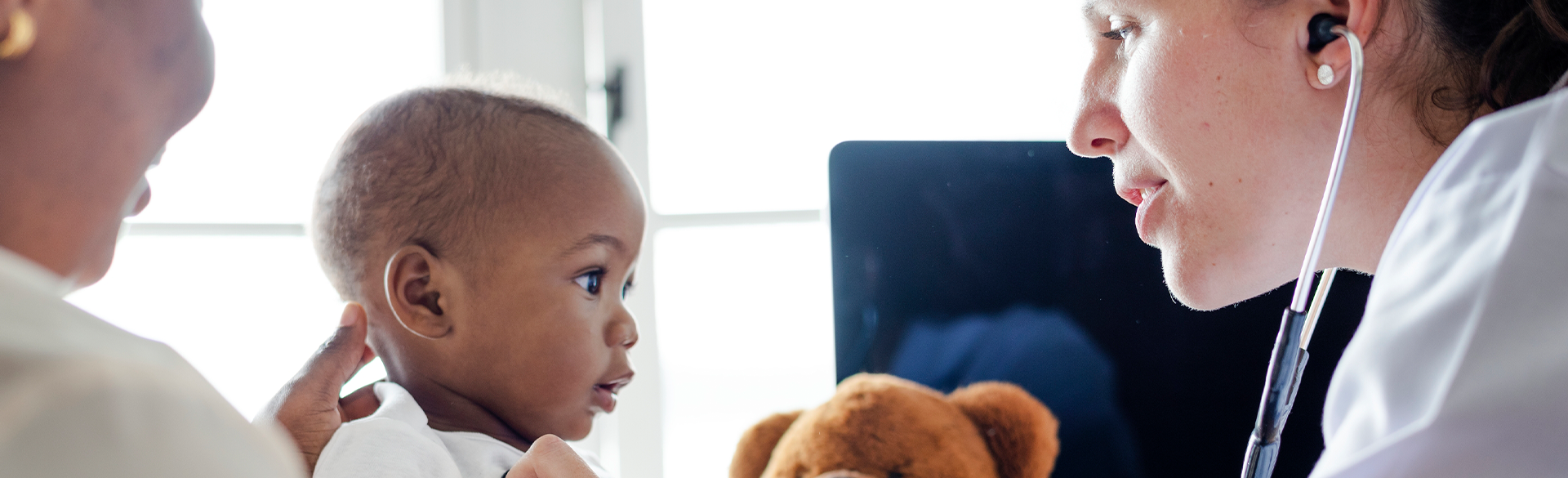
CCTSI Research Seeks to Establish Safe Oral Antibiotic Doses for Infants
4/8/2025New research at the Colorado Clinical and Translational Sciences Institute (CCTSI) aims to establish safe oral antibiotic doses for infants, potentially reducing hospital stays and improving care for newborns with infections....Read More
-

Forty Years of Leading High-Impact Research
3/3/2025Wendy Kohrt, PhD, is stepping down from her role as CCTSI's Associate Director of Resources and Services after nearly 40 years....Read More
-
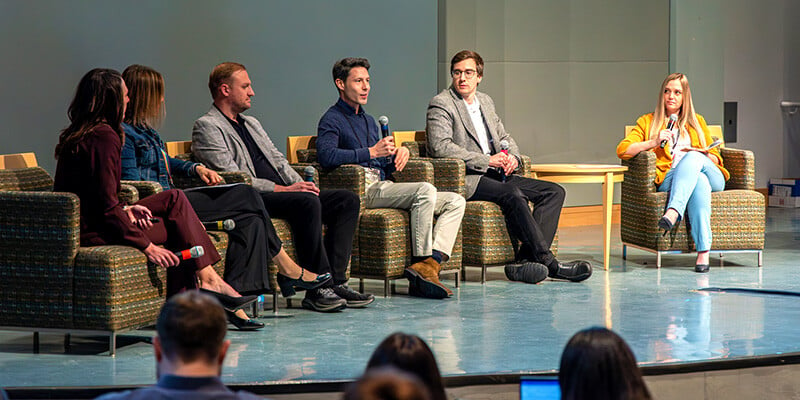

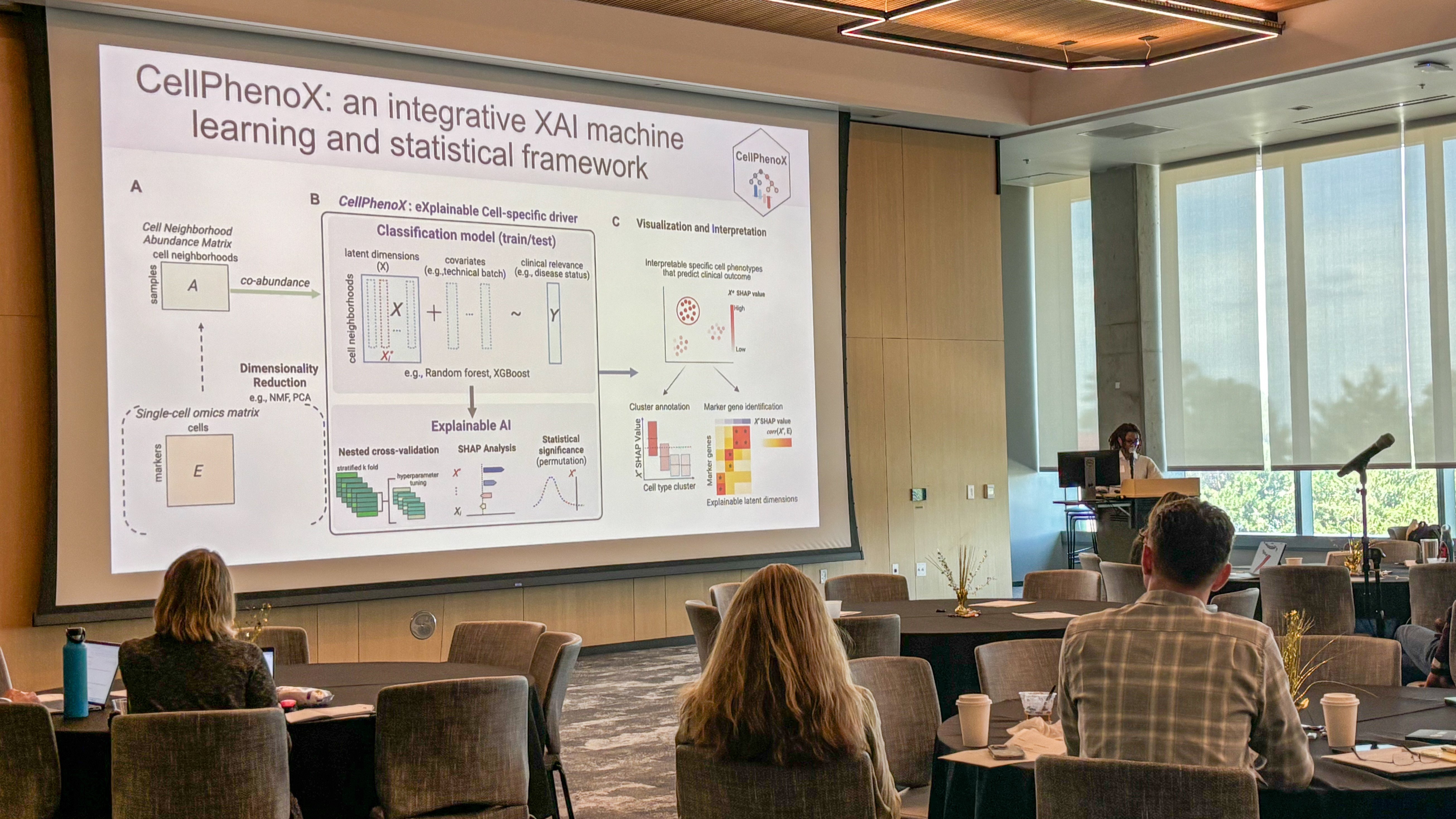
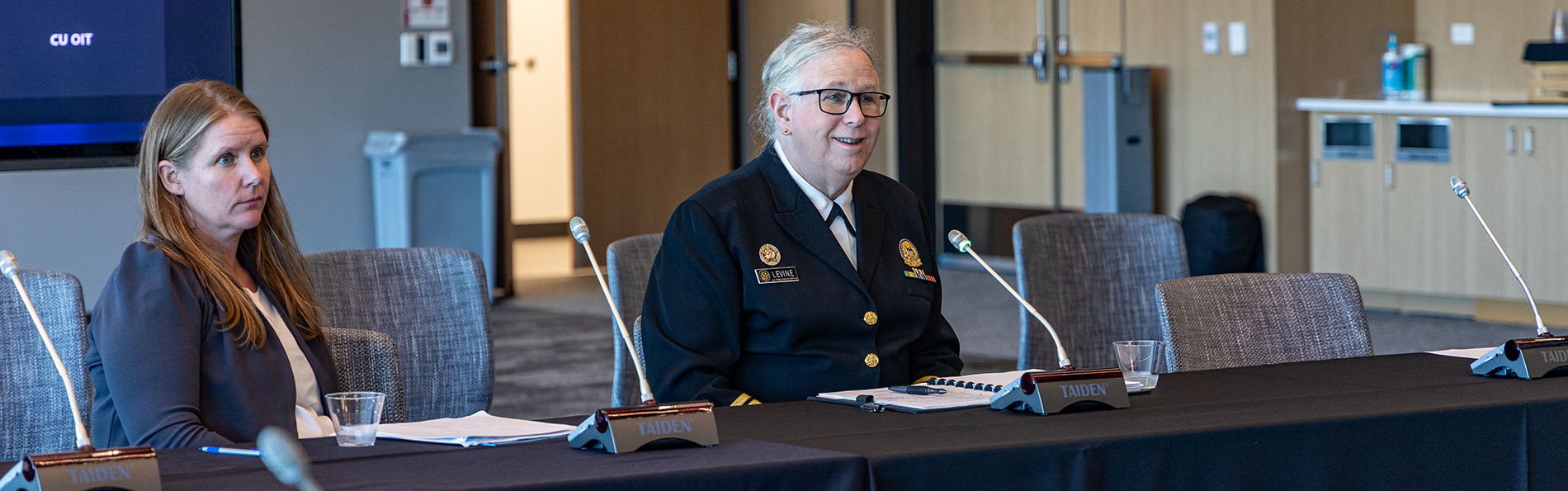
CU Anschutz Readies for AI's Evolution
2/24/2025CU Anschutz gathered to consider how, as Chancellor Don Elliman said, to unlock the promise of AI while at the same time minimizing the potential dangers....Read More
ACCELERATING RESEARCH TO IMPROVE HEALTH
The CCTSI helps build research teams of the future, speed the development of new treatments and improve human health

Resources and Services
CCTSI supports translational research by providing resources, integrating facilities and services, developing innovative research tools and catalyzing interdisciplinary research programs.Funding
Each year, the CCTSI awards more than $2 million in pilot grants, microgrants, and pre-doctoral and post-doctoral research scholar awards. You can find more information on each grant and how to apply
Education, Training and Career Development
The Education, Training and Career Development (ETCD) program provides clinical-translational scientists and trainees with knowledge, training and career skillsClinical Translational Research Centers
The CTRC provides infrastructure, resources and dedicated space to conduct clinical research.

















%20(1).png)


-1.png)









.jpg)






.png)