University of Colorado Department of Pathology
University of Colorado Department of Pathology
University of Colorado Department of Pathology
University of Colorado Department of Pathology
University of Colorado Department of Pathology
Welcome to the Department of Pathology
University of Colorado, School of Medicine
Welcome to the Department of Pathology of the University of Colorado, Anschutz Medical Campus. The Department has grown substantially in the past 15 years, from 40 to 120 faculty in parallel with the remarkable growth of our hospital based affiliates as well as the city and county of Denver. Our work is value driven and focused on scientific investigation, lifelong learning and a balance of personal and professional values. In addition to a vibrant and highly competitive residency program with 25 positions, we offer 9 fellowships and participate in numerous graduate school and the MD/PhD program of the CU School of Medicine. Our faculty and staff are diverse and gender balanced, with women comprising 50% of our residents, Assistant, Associate and Full Professors (including those with tenure).
Continue to full message

Academic Programs
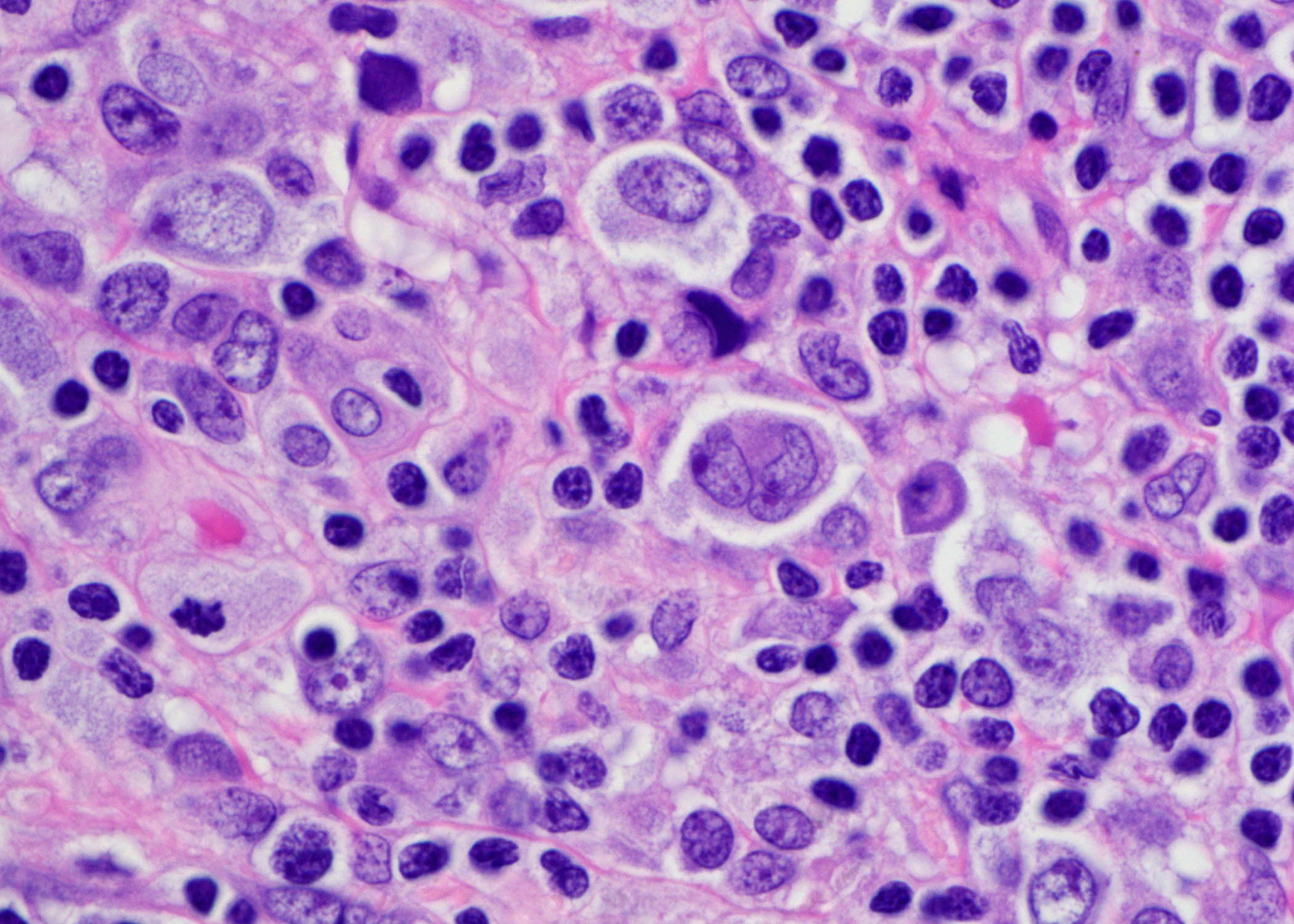
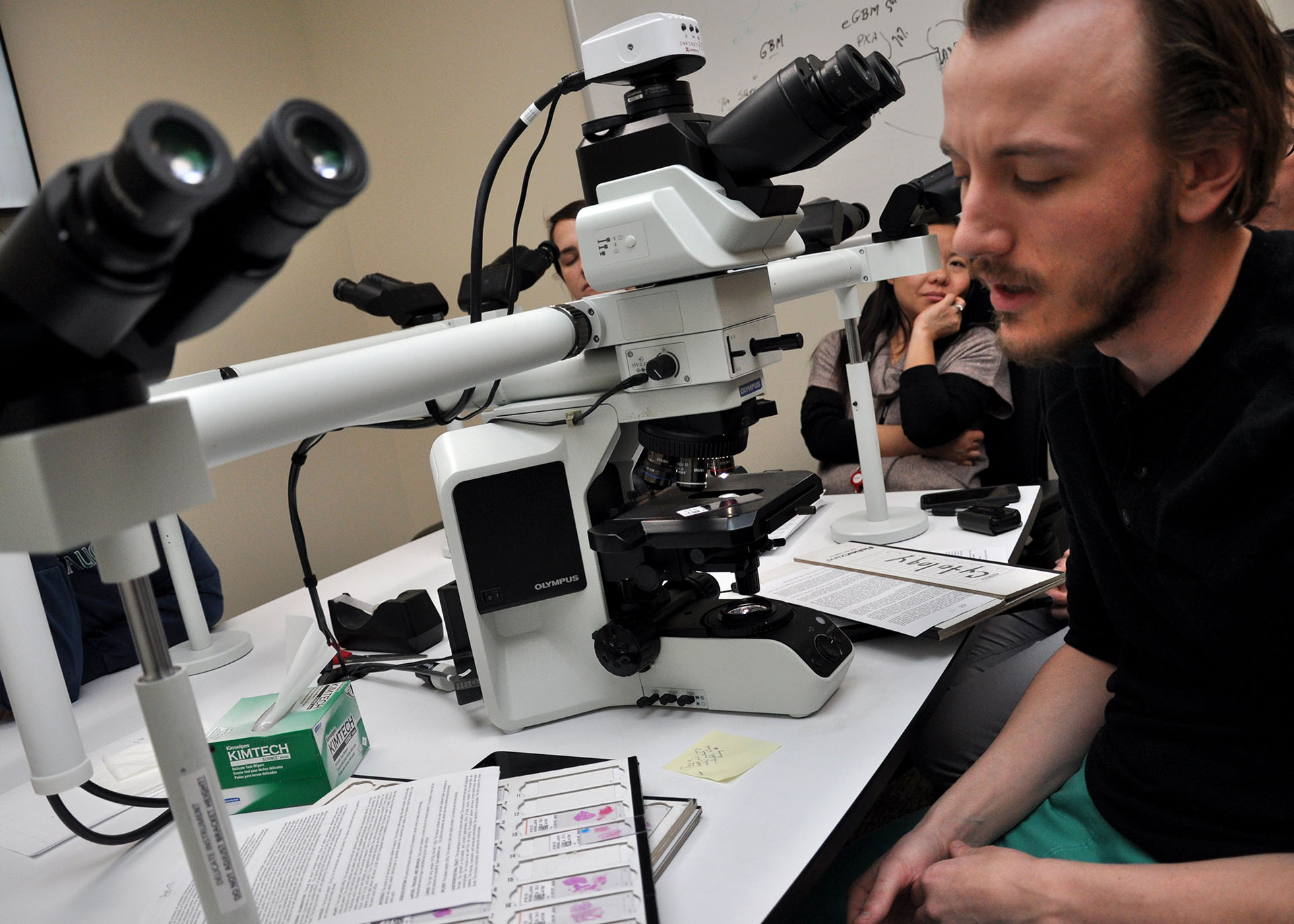
Anatomic Pathology
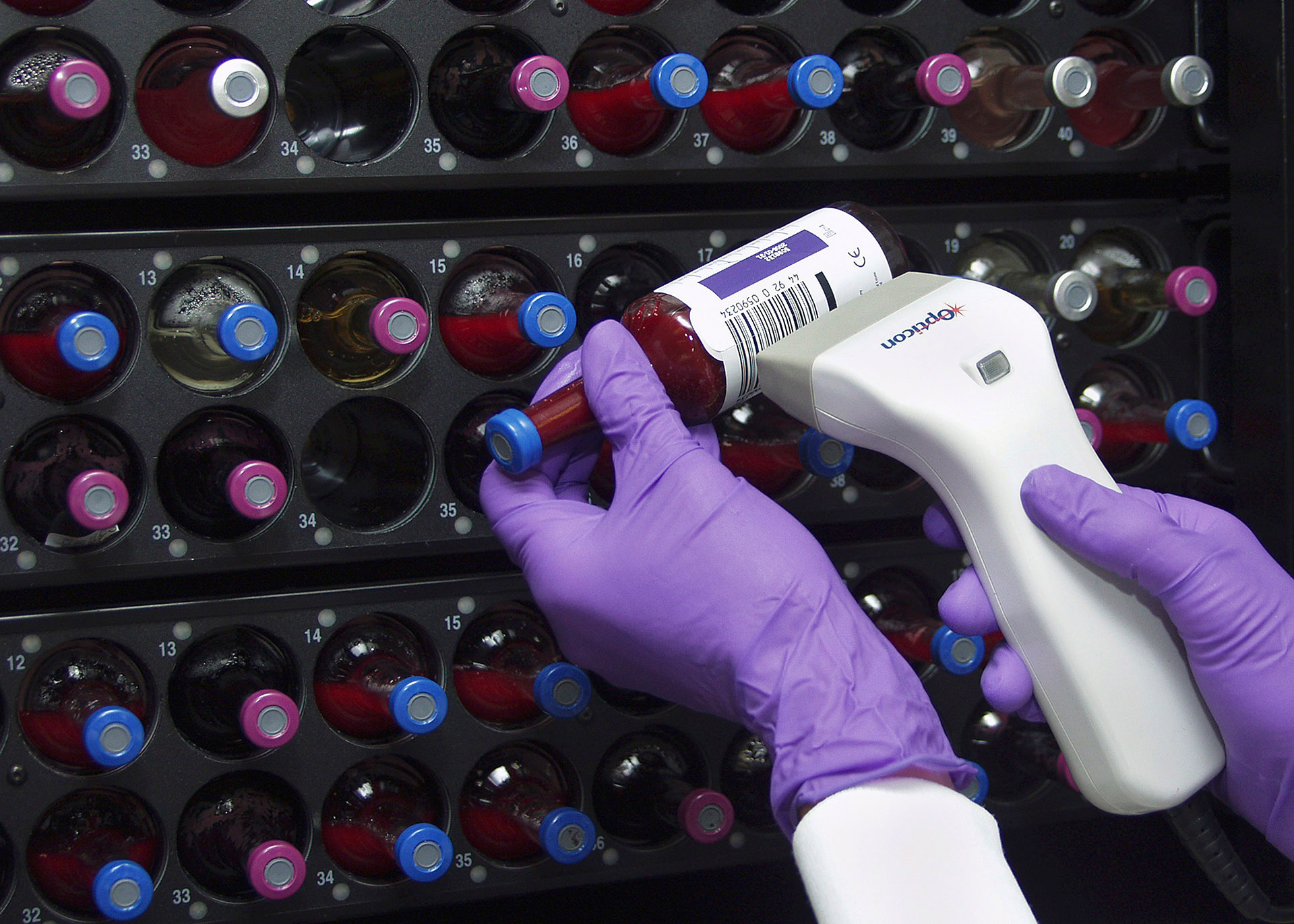

Clinical Pathology
- Clinical Laboratory
- Research
- Rotations
- Fellowships
Residency Program Applicants
We invite you to use the links below for information on our Residency Program!
Department of Pathology Residency Information Videos
Department of Pathology Academic Programs

Ann D. Thor, M.D.
Professor and Todd Chair of Pathology